Wolfram Data Repository
Immediate Computable Access to Curated Contributed Data
Categorization of the pathogenicity data of human missense variants
| "Chromosome" | string including "Chromosome" and a number from 1 to 22 or a letter M,X,or Y. |
| "RawDataSize" | total rows for each chromosome in the original database |
| "TotalPositions" | total positions for each chromosome |
| "TotalMutations" | total missense mutations for each chromosome |
| "Genome" | the genome build |
| "TotalUniprotID" | total uniprot IDs |
| "TotalTranscriptID" | total transcript IDs |
| "TotalAminoAcidVariations" | total amino acid changes |
| "PathogenicityQuartiles" | pathogenicity Quartiles 1/4, 2/4, 3/4 |
| "PathogenicityMean" | pathogenicity Mean |
| "PathogenicityLikelyBenign" | percentage of "likely_benign" classifications |
| "PathogenicityLikelyPathogenic" | percentage of "likely_pathogenic" classifications |
| "PathogenicityAmbiguous" | percentage of "ambiguous" classifications |
| "Chromosome" | string including "Chromosome" and a number from 1 to 22 or a letter M,X,or Y. |
| "Position" | genome position (1-based) |
| "ReferenceNucleotide" | reference nucleotide (GRCh38.p13 for hg38) |
| "AlternativeNucleotide" | alternative nucleotide |
| "Genome" | genome build |
| "UniprotID" | UniProtKB accession number of the protein in which the variant induces a single amino-acid substitution (UniProt release 2021_02) |
| "TranscriptID" | Ensembl transcript ID from GENCODE V32 (hg38) |
| "AminoAcidVariation" | Amino acid change induced by the alternative allele,in the format: Reference aminoacid-POS_aa-Alternative amino acid |
| "Pathogenicity" | predicted probability of a variant being clinically pathogenic |
| "Classification" | derived using the following thresholds: "likely_benign" for Pathogenicity < 0.34; "likely_pathogenic" for Pathogenicity > 0.564; and "ambiguous" otherwise |
| "ExternalIdentifier" | ExternalIdentifier of the protein |
| "Name" | common name of the protein |
| "Sequence" | amino acid sequence of the protein |
| "Mutations" | all possible mutations in the protein |
| "Score" | AlphaMissense pathogenicity score by mutations of a protein |
| "Status" | status of pathogenicity (B: Likely benign, A: Ambiguous, P: Likely pathogenic) |
| "MeanPathogenicity" | Mean pathogenicity per residue of a protein |
| "MedianPathogenicity" | Median pathogenicity per residue of a protein |
| In[1]:= |
| Out[1]= | 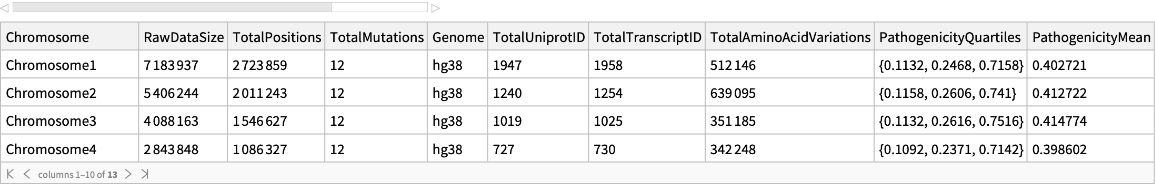 |
Retrieve the data for ChromosomeX:
| In[2]:= |
| Out[3]= | 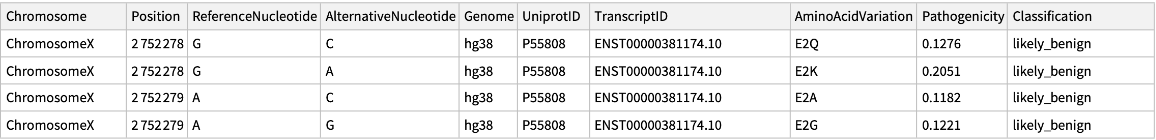 |
Get a random sample of the Positions in ChromosomeX:
| In[4]:= |
| Out[4]= |
Get a random sample of the transcriptIDs in ChromosomeX:
| In[5]:= |
| Out[5]= |
Get a distribution of the ChromosomeX pathogenicity level for all possible missense variations:
| In[6]:= | ![BoxWhiskerChart[chrX[All, #Pathogenicity &], "Mean", PlotLabel -> "Pathogenicy level distribution", FrameLabel -> {"ChromosomeX", "Pathogenicity level"}]](https://www.wolframcloud.com/obj/resourcesystem/images/225/225dffa1-1f90-49d1-912c-d751dd984043/18672d26a5862b04.png) |
| Out[6]= | 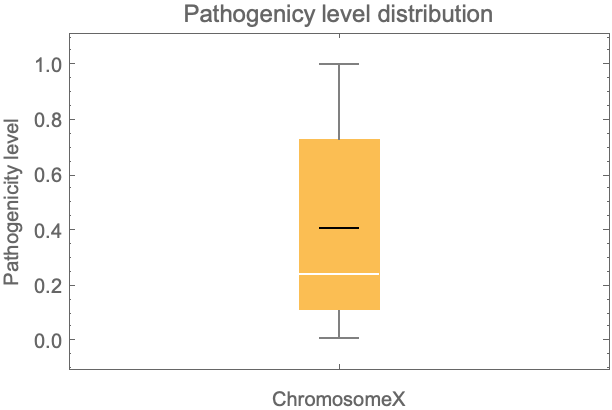 |
Get a summary of the classification of the ChromosomeX pathogenicity level for all possible missense variations:
| In[7]:= |
| Out[7]= | 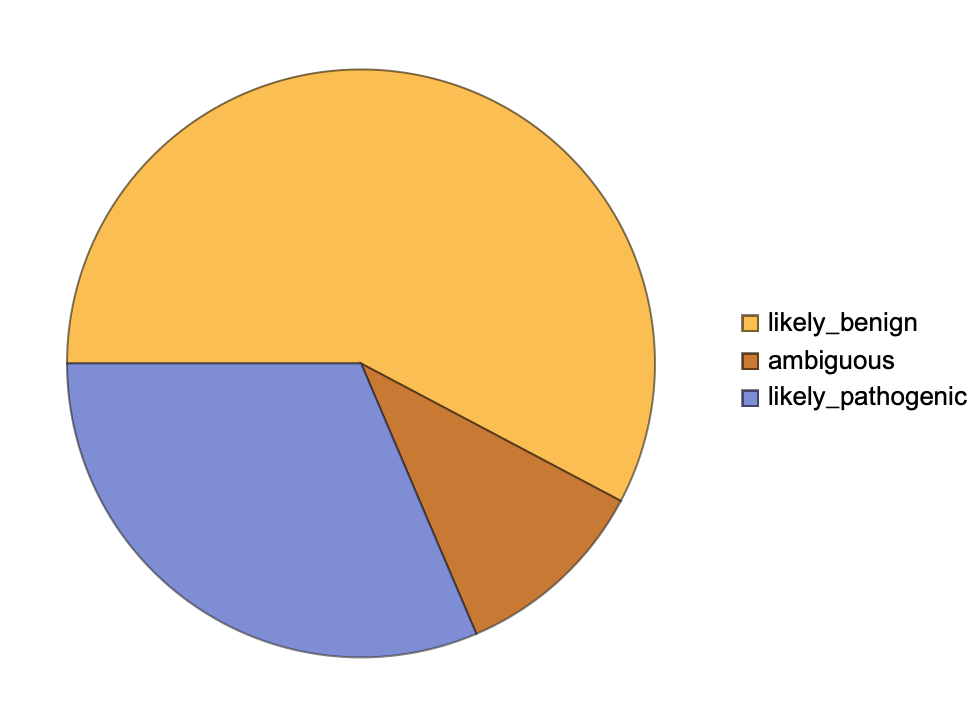 |
| In[8]:= |
| Out[8]= |
Select a protein from the set:
| In[9]:= |
| Out[9]= |
Find the protein's properties:
| In[10]:= |
| Out[10]= | 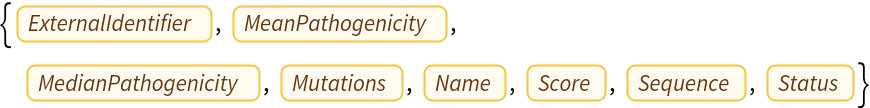 |
Get the protein's name:
| In[11]:= |
| Out[11]= |
Get the protein's amino acid sequence:
| In[12]:= |
| Out[12]= |  |
Visualize the mean and median AlphaMissense pathogenicity per residue:
| In[13]:= |
| Out[13]= | 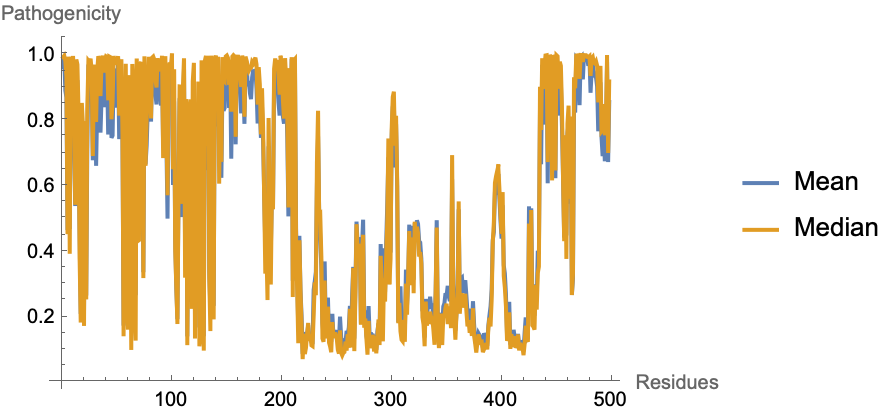 |
Get the AlphaMissense pathogenicity score for a specific residue:
| In[14]:= |
| Out[15]= |  |
Get the AlphaMissense pathogenicity status for a specific residue:
| In[16]:= | ![pathAssoc = <|"B" -> "Likely benign", "A" -> "Ambiguous", "P" -> "Likely pathogenic"|>;
AssociationThread[protein["Mutations"][[resNumber]], Map[pathAssoc, Characters[protein["Status"][[resNumber]]]]]](https://www.wolframcloud.com/obj/resourcesystem/images/225/225dffa1-1f90-49d1-912c-d751dd984043/5ca367e8ba8d8c61.png) |
| Out[17]= |  |
Get the name of a specific protein:
| In[18]:= |
| Out[18]= |
Get the pathogenicity information associated to ChromosomeX in position 71765227:
| In[19]:= | ![PathogenicityData[chromosome_String, mutationPosition_Integer, "Pathogenicity"] := Block[{rawData0}, rawData0 = ResourceData[\!\(\*
TagBox["\"\<Alpha Missense\>\"",
#& ,
BoxID -> "ResourceTag-Alpha Missense-Input",
AutoDelete->True]\), chromosome][
Select[MatchQ[#[[2]], mutationPosition] &]]; Dataset[Association[{"Position" -> #[[2]], "Mutation" -> {#[[3]] -> #[[4]]}, "TranscriptID" -> #[[7]], "AminoAcidVariation" -> #[[8]], "Pathogenicity" -> #[[9]], "Classification" -> #[[10]]}] & /@ Normal[rawData0]]]](https://www.wolframcloud.com/obj/resourcesystem/images/225/225dffa1-1f90-49d1-912c-d751dd984043/06e8faf2184617ac.png) |
| In[20]:= |
| Out[20]= | 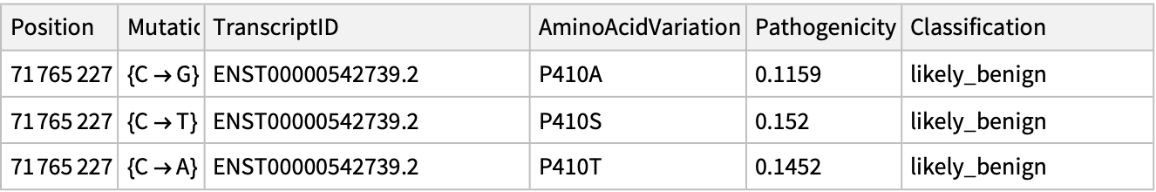 |
Get the pathogenicity information associated to ChromosomeX for a list of positions:
| In[21]:= | ![PathogenicityData[chromosome_String, mutationPosition_List, "Pathogenicity"] := Block[{rawData0}, rawData0 = ResourceData[\!\(\*
TagBox["\"\<Alpha Missense\>\"",
#& ,
BoxID -> "ResourceTag-Alpha Missense-Input",
AutoDelete->True]\), chromosome][
Select[MemberQ[mutationPosition, #[[2]]] &]]; Dataset[Association[{"Position" -> #[[2]], "Mutation" -> {#[[3]] -> #[[4]]}, "TranscriptID" -> #[[7]], "AminoAcidVariation" -> #[[8]], "Pathogenicity" -> #[[9]], "Classification" -> #[[10]]}] & /@ Normal[rawData0]]]](https://www.wolframcloud.com/obj/resourcesystem/images/225/225dffa1-1f90-49d1-912c-d751dd984043/7fecc90d0859bbec.png) |
| In[22]:= |
| Out[22]= | 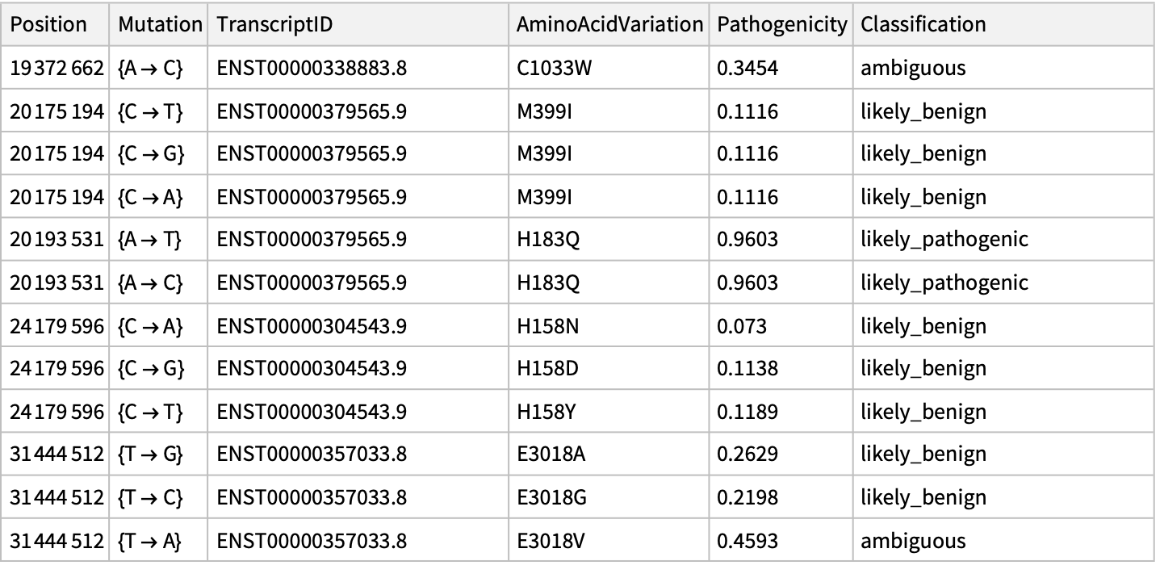 |
Get the pathogenicity distribution associated to all the positions of all the chromosomes:
| In[23]:= | ![data = Table[Normal[ResourceData[\!\(\*
TagBox["\"\<Alpha Missense\>\"",
#& ,
BoxID -> "ResourceTag-Alpha Missense-Input",
AutoDelete->True]\), "Chromosome" <> ToString[i]][
All, #Pathogenicity &]], {i, {Sequence @@ Range[1, 22], "M", "X",
"Y"}}];](https://www.wolframcloud.com/obj/resourcesystem/images/225/225dffa1-1f90-49d1-912c-d751dd984043/7644c05ec695f98f.png) |
| In[24]:= | ![BoxWhiskerChart[data, "Mean", ChartLabels -> {Sequence @@ Table["chr" <> ToString[i], {i, 1, 22, 1}], "chrM", "chrX", "chrY"}, PlotLabel -> "Pathogenicy level distribution", FrameLabel -> {"", "Pathogenicity level"}]](https://www.wolframcloud.com/obj/resourcesystem/images/225/225dffa1-1f90-49d1-912c-d751dd984043/4087ee817266c1c8.png) |
| Out[24]= | 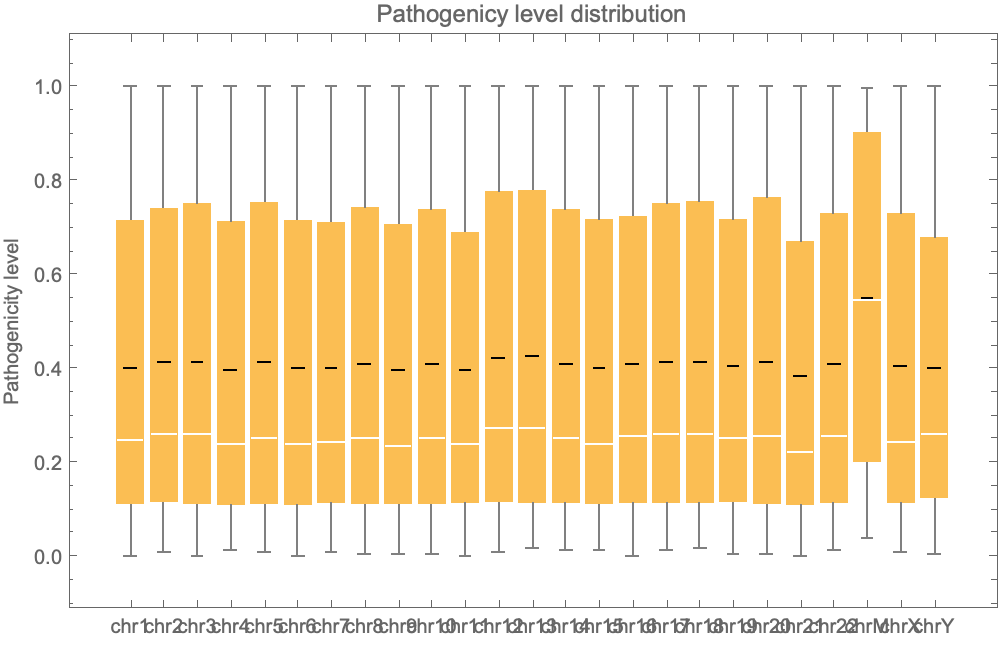 |
Wolfram Research, "Alpha Missense" from the Wolfram Data Repository (2024)